To purpose of this research project was to understand how season influences the taxonomic and metabolic composition of microbial communities inhabiting three Finger Lakes in central New York, and to understand how future climate scenarios will affect these characteristics. To accomplish these goals, two major activities were performed. First, we performed a year-long time series analyses of microbial community gene expression in concert with high resolution measurements of ambient temperature, salinity, oxygen, and overall productivity (via measurement of phytoplankton chlorophyll a concentration). Next, we performed a series of 3 experiments where microbial communities were enriched over ambient CO2 conditions (thus lowering pH) and raised temperature to simulate lake water conditions in the future. Our initial idea was to examine in these latter experiments the dynamics of functional genes or taxonomic constituents which demonstrated seasonality and which may have been sensitive to future climate scenarios.
All three lakes demonstrated a seasonal pattern of temperature, with greatest values in July and lowest values in February (Fig. 1). Amongst the three lakes, Owasco had the greatest variation in annual temperature from 24.4oC in mid-summer and 0.5oC in February, while Cayuga had the least variation from 23.1oC in mid-summer and 2oC in January. The overall change in temperature reflects polymictic conditions throughout the year, although this may also be confounded by the sampling location in very shallow water.
In contrast to Temperature, chlorophyll a concentration (which indicates overall phytoplankton biomass) showed highest values in Spring, although the timing of maximum phytoplankton concentration differed between lakes. Owasco experienced a Spring Bloom in April, while both Seneca and Cayuga Lakes experienced Blooms in May – June. A pronounced ‘clear water phase’ after bloom exhaustion occurred in Owasco in May, however was absent in Seneca and Cayuga. Bacterial abundance generally increased across all three lakes from December through mid-Summer, with highest summer abundances occurring in Owasco, and lowest in Seneca.
Metatranscriptome libraries (n = 30) prepared from the three lakes generated a total of 14,008,885 annotated sequences, representing ~ 5% of total sequence reads overall. The remaining sequence reads did not match any sequences in NCBI GenBank.
Amongst sequence reads, Bacteria comprised the largest proportion of reads, with fewer sequences from Eukaryotes, and only small numbers from Archaea or Viruses (Fig. 2).
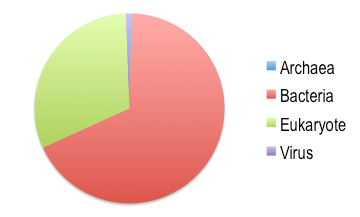
Fig. 2: Phylogenetic distribution of metatranscriptomic reads in the Finger Lakes.
Amongst functional annotations, transcripts were dominated by genes involved in central metabolism, predominately Carbohydrates, Protein Metabolism, Amino Acids and Derivatives, Photosynthesis and Respiration and Vitamin metabolism (Fig. 3). There were fewer genes involved in predicted biogeochemical cycles, including Nitrogen Metabolism, Phosphorus Metabolism, and Iron Cycling. There were few transcript or even transcript families (i.e. genes sharing similar function) which were shared between sampling dates. Temporal patterns of photosynthesis, respiration and key biogeochemical cycling genes did not demonstrate predicted patterns over season (Fig. 4). However, virulence and disease-related genes demonstrated a hump-shaped distribution over the time series sampling, with greater transcripts related to virulence in winter months compared to fall or spring.
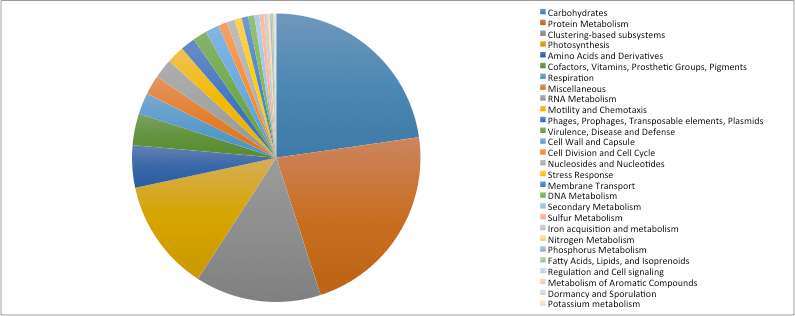
Fig. 3: Functional annotations of transcripts from the Finger Lakes.

Fig. 4: Percentage of gene transcript reads corresponding to key biogeochemical cycles (top and mid panels) and virulence genes (bottom panel).
Clustering based on functional categories of transcripts did not reveal distinct patterns with lakes, season, or between replicate metatranscriptomes from the same sampling time and lake (Fig. 5).

Fig. 5: Multiple dimensional scaling of metatranscriptomes based upon similarity between transcript profiles of functional annotation.
The abundance of viral related transcripts demonstrated a strong seasonal pattern, with much greater abundance in winter months than in fall or spring. The abundance of viral transcripts was over 1000X greater in cooler months (Fig. 6)
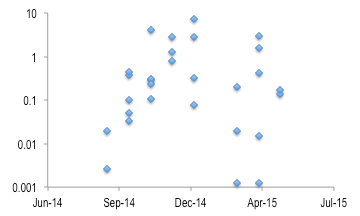
Fig. 6: Relative percent of viral-related transcripts in metatranscriptomes over the time series sampling.
Identification of RNA Viral Phylotypes in Metatranscriptomes: Based on our observation of viral related gene variation over seasonal cycles, and the season cycling of virulence-related genes, we focused on RNA viruses for further analyses. After sequence assembly, many near-complete RNA viral genomes were obtained. Recruitment of reads against RNA viral genomes identified five genomes with large (>10X) coverage and which were represented in >half of all metatranscriptomes. These five contigs, named Pico-246, Pico-248, Noro, Orbi and Dicistro were selected for further analyses.
Pico-246 is most similar to Beihai Papia Shell Virus (APG78606.1) sharing 45% identity (E-val 8e-157). Pico-248 is most similar to Behai picorna-virus like, (APG78024.1), sharing 44% identity (E-val 0), the host of which is a sesarmid crab. Noro is most similar to Wenling Crustacean Virus (APG78478.1), sharing 44% identity (E-val 2e-31). Orbi is most similar to the mosquito Orungo Virus (AFX73387.1), sharing 30% identity (E-val 4e-17). Finally, Dicistro is most similar to Bee Paralysis virus (YP_003622540.1), sharing 24% identity (E-val 5e-76). Taken together, these viruses are likely mollusk (Pico-246) and aquatic arthropod (all but Pico-246) viruses that are common constituents of the Finger Lakes.
Temporal Quantification of RNA Viral Phylotypes: Quantitative PCR primers and probes were developed around all 5 viral genotypes, targeting capsid protein motifs. The qPCR primers/probes were then applied against the 5um size fraction extracts from the time series samples. Our rationale for using the larger size fraction was that the viruses detected in the metatranscriptomes were likely associated with tissue or free cells (i.e. environmental DNA or eDNA) and would more likely be associated with the larger size fraction.
The five viral phylotypes demonstrated different temporal patterns with season and between lakes. Pico-246 had greatest abundance during winter months in Owasco Lake, but had a very large abundance in Seneca and Cayuga during the spring (Mar-April 2015). Pico-248 was absent entirely from Seneca Lake and very small abundances in Cayuga Lake, but had a large increase in winter months in Owasco Lake. Noro and Orbi both had no clear pattern with season in any lake. Dicistro was absent from Cayuga Lake, but had large swings in abundance in Seneca Lake and Owasco Lake during Spring and Winter, respectively.
Pairwise correlation between viral phylotype abundance, temperature, chlorophyll and bacterial abundance revealed few significant (p<0.05) correlations. When considered as a single data set (i.e. combining all lakes together), there were no significant correlations between any viral phylotype and environmental conditions, however Dicistro correlated with Pico-246 (R2 = 0.67). Considering each lake separately, however, revealed that Pico246 correlated with chlorophyll a concentration (R2 = 0.34) in Cayuga Lake, with temperature (R2 = 0.37) in Owasco Lake, and with Dicisto (R2 = 0.62) in Seneca Lake, and with Noro (R2=0.44) in Cayuga Lake. Additionally, Noro correlated with Orbi (R2 = 0.43) and with Chlorophyll a concentration (R2 = 0.59) in Owasco Lake.
Experimental Incubations: The addition of acidified water (in Oct 2014, pH 5.4, in Jan 2015, pH 4.84 and in Apr 2015, pH 5.03) to ambient lakewater in mesocosms (pH 7.63, 7.54 and 7.72 for the three experiments, respectively) resulted in decreased pH in mesocosms by pH 1.28 – 1.76. Temperature in treated mesocosms was also raised relative to controls by 2.8 – 8.9 degrees C by aquarium thermostats. Temperature rise was greatest in January mesocosms when ambient water temperatures were ~0.5oC, and least in summer when ambient water temperature was 24.2oC.
The dynamics of five viral phylotypes revealed variable responses depending on phylotype, lake and date (Fig. 8). Pico-248 was largely absent from mesocosm experiments. Pico-246 was absent in October 2014, but in January and April 2015 was present across most experiments. Orbi was present mostly in January 2015 experiments, but was also present in the Seneca Lake experiments in April 2015. Dicistro was mainly present in Seneca Lake, while Noro was represented only in Owasco Lake in January and April 2015.
Acidification and temperature did not have a consistent impact on the abundance of any viral phylotype, while containment alone did impact overall viral abundance (Fig. 8). For example, treatment mesocosms bore detectable phylotypes, where controls and/or initial phylotype abundances were absent in 3 experiments (representing Pico-248, Orbi and Dicistro). Conversely, initial and/or control treatments had detectable viral copies but treatment were absent in 9 experiments, representing Orbi (4 experiments), Dicistro (2 experiments), Pico-248 (2 experiments) and Noro (1 experiment). Amongst experiments where a statistical comparison could be made between detectable control and treated incubations (n=14), there were few (n= 3) significant differences. Noro virus was significantly higher in treated incubations relative to controls and initial abundances in Owasco Lake in January 2014, but was significantly lower than both controls and initial abundances in the same lake in April 2015. Dicistro also had lower abudnances in Cayuga Lake in October 2014 than in controls and initial values.

Fig. 6: Abundance of viral phylotypes in mesocosm incubation experiments in Owasco (A-C), Seneca (E-G) and Cayuga (H-J) Lakes. Experiments occurred in October 2014 (A,E,H), January 2015 (B,F,I) and April 2015 (C,G,J). The abundance of phylotypes (copies per L) was determined by qRT-PCR based on the 5um size fraction.
Discussion
The pattern of gene expression in bacterioplankton were mostly similar to those reported in previous studies of geographic, depth-related and diel changes in gene expression (Hewson, Poretsky et al. 2009, Hewson, Poretsky et al. 2009, Poretsky, Hewson et al. 2009, Hewson, Poretsky et al. 2010). While the majority of transcripts across all libraries were associated with bacteria, there were also a large number of eukaryotic transcripts present, which may represent picoeukaryotic organisms that passed through the 5um pre-filters used in this study. The very low percentage of total archaeal reads was surprising, since these typically comprise ~10 – 20% of surface water bacterioplankton cells in marine waters. These results point to the lower contribution of archaea relative to bacteria in freshwater ecosystems relative to other aquatic habitats. Functional gene categories were also consistent with previous work showing core metabolic genes dominating overall gene expression. For example, previous studies have found a large proportion of transcripts overall are associated with carbohydrate and protein metabolism, RNA and DNA metabolism, and the metabolism of Amino Acids and Derivatives, which in this study comprised > 60% of all transcripts. The presence of large numbers of photosynthesis and respiration-related genes is also consistent with surface water productivity. Previous work in central ocean gyres have found that photosynthesis-related genes, which have high turnover and short half-lives, can account for up to 22% of total gene transcripts in any microbial transcript pool.
The seasonal pattern of gene expression had surprisingly little seasonal variation. Photosynthesis and respiration-related genes had a slight decrease in relative representation in mid-winter, however was constant for the remainder of the year. Similarly, important biogeochemically-related genes involved in S, P and N cycles were constant and variable over the year. A slight increase in P-metabolic genes coincided with the Spring Bloom of 2015. The lack of change in N-metabolic genes may at this time may indicate P-limitation of lake waters which was alleviated by melting ice and release of P from sediments. Interestingly, virulence-related genes (i.e. those that enable pathogens to cause disease) increased over 100 fold between Fall 2014 and Spring 2015, with greatest values in March 2015 (coinciding with ice-out on all lakes), suggesting a higher prevalence of disease-causing organisms as lake waters warm. Clustering of metatranscriptomes based on shared gene families revealed no pattern between lakes, between sampling dates, or even between replicates. This result is consistent with the lack of change in expressed gene families over the seasonal cycle, and high variation in relative transcript frequency between gene families between replicate samples.
Taken together, the overall transcriptional profile indicated a photosynthetically active community of microorganisms that was highly variable between water samples, but was also remarkably invariant in metabolism over the seasonal cycles. The major difference in community transcripts pre- and post-ice out on the lakes appeared related to the expression of virulence-related genes which may because of several factors. First, new pathogens may be introduced to lake water by inflow of meltwater from the catchment. Second, non-pathogenic allochthonous microorganisms may bear more similarity to cultivated pathogens in clinical settings, and thus may be more annotated as pathogens than the relatively large number of uncultivated aquatic bacteria. Finally, there may be proliferation of pathogens in aquatic habitats during ice-out since the overall abundance of organisms may increase as spring progresses, which facilitates transmission of pathogens between uninfected and infected organisms. Bacterial abundance and chlorophyll a concentration both demonstrated strong increases from winter to spring 2015.
Because we observed the seasonal trend of increasing virulence genes from Winter into Spring, we further looked at the abundance of viral-related transcripts over the time series. The abundance of viral transcripts (which represent expressed genes of DNA viruses, genomes of RNA viruses) had a U-shaped distribution, with lower relative abundances in Fall and Spring compared to mid-winter. These transcripts included predominately bacteriophages, but RNA viruses were also well represented in their pool. Bacteriophage may increase as abundance of hosts decrease in winter due to induction of the lytic cycle of lysogens, or this may also represent increased susceptibility of hosts as they scavenge for nutrients in the lake by loading additional transmembrane proteins (and, consequently, surface cell receptors) for nutrient uptake.
The observed lack of change in cellular microbial transcripts over seasonal cycles, but increase in virulence-related genes during spring progression, along with hump-shaped distribution of virus-related reads peaking in winter led us to further investigate viral phylotypes over the course of the experiment. Examination of metatranscriptomes revealed several well covered and complete/near-complete genomes of RNA viruses, mainly Picornaviruses (+ve sense ssRNA viruses). This group of viruses is known to infect a wide range of hosts including eukaryotic microorganisms (e.g. photosynthetic flagellates), invertebrates and vertebrates. We selected 5 RNA viral phylotypes based on their wide representation between metatranscriptomes, their coverage in each metatranscriptomes, and also because they may infect ecologically important constituents of the lake ecosystem. Initial annotation in May 2015 indicated that these corresponded to pathogens of waterfowl (Duck Hepatitis Virus – Noro), plants (A pathogen of Oryza – Orbi), an insect (Bee paralysis virus – Dicistro), and two that were distantly related to viruses infecting other arthropods (Pico-248 and Pico-246). An extensive sequencing study in Mid-2015 (Shi, Lin et al. 2016) elucidated a wide range of new Picornaviruses isolated from diverse hosts which caused reclassification of those viruses targeted in our study. Based on these newly available sequences, Noro appears more closely related to a shrimp virus (and more loosely related to other invertebrate picornaviruses than to any eukaryotic microbial or vertebrate virus), Orbi is more closely related to a Mosquito virus, Pico-246 to a Mollusk virus, and Pico 248 most similar to another crustacean virus. Dicistro remains most similar to the Bee Paralysis virus, which is implicated in colony collapse disorder in apiaries.
Because of the putative origin of these viruses (invertebrates) we chose to focus our work on the >5um size fraction, since few cells of invertebrates are expected in the bacterioplankton (5 – 0.2 um) size fraction. Quantitative PCR of the 5 viral phylotypes revealed that viral phylotypes either had inconsistent pattern with season (Orbi and Noro), or maxima in warmer months (Dicistro and Pico-246), in cooler months (Pico-248). These patterns may reflect host metabolism or inflow of allochthonous viruses. Both Orbi and Noro may be pathogens of zooplankton which are present year-round in the lake and which may experience sporadic mortality events by viruses. Pico-248 may be a pathogen of a zooplankton species which may experience greater mortality (and greater transmission) as abundances increase in Spring relative to winter, when their abundances are lower (e.g. Daphnia or copepods). The greater abundance of Dicistro in spring may indicate that it is an allochthonous virus which may come into lake ecosystems during ice-out. Pico-246, which may infect Quagga or Zebra mussels in the lake, may have higher abundances in Spring relative to winter due to enhanced productivity and transmission on vectors (e.g. particles), since its mortality is likely more density independent than pelagic organisms. To the best of our knowledge, this is the first seasonal report of viruses possibly infecting invertebrate plankton in lake plankton, and provides important information on their potential role in lake habitats.
The intention of the experimental incubations was to simulate future climate scenarios and understand how viruses may respond to these conditions. In reality, our treatment (temperature and acidification) was significantly greater than that expected in the next 100 years. However, by strongly changing pH and temperature, we can understand how viruses may respond on short time scales, since subtler changes in physic-chemical conditions would be expected to elicit subtler changes. We observed very few statistically significant patterns of viral abundance relative to treatment, with most viruses being either completely absent, or being detectable in only treatment (but not in initial or controls) or in initial and/or controls but not in treatment. We can interpret these experiments in two ways. First, those viruses that were present in only treatment mesocosms but absent in both initial and controls may represent viruses that are induced by the conditions under which they are imposed. Second, those viruses that are absent in treatments but present in both initial and controls may be negatively impacted by the treatment. Orbi and Dicistro both were negatively and positively impacted by treatment (by this criterion) in different experiments, but were more negatively affected than positively. This may indicate that Orbi and Dicistro, which are putatively allochthonous viruses, may be instable under lower pH and higher temperature conditions once they enter the lake habitat. Similarly, Pico-248 had an equal number of positive and negative effects of treatment. A better comparison of treated vs control incubations comes from experiments where we detected viral copies in both conditions. Of these, Noro had conflicting results, with both positive and negative impacts of treatment occurring in adjacent months. Dicistro had significantly lower abundance in treated experiments than controls.
Taken together, our results suggest that picornaviruses are largely unaffected by enhanced temperature and lower pH conditions, even then these are many times more than expected in the next 100 years with climate change. The only viral phylotype which was significantly affected was Dicistro, which in most cases was significantly and negatively impacted by the imposed conditions. Because this may be an allochthonous virus, it may simply have more unstable particles in harsher water conditions and may therefore be less likely to be reintroduced by insects consuming lake water, or in crops irrigated with lake water, in the future.
The study revealed that:
- The abundance of pathogen-related genes increased during spring months relative to winter and fall, and that viral transcripts were greatest in winter;
- Most cellular microbial transcripts were relatively invariant over season, even those relating to productivity, like photosynthesis and respiration;
- Picornaviruses possibly infecting invertebrates in the lake ecosystem, or infecting allochthonous insects, have variable seasonal pattern, with some experiencing strong swings relating to spring bloom phenomena, while others having sporadic increases in abundance;
- Future climate scenarios will not impact the abundance of most (and putatively autochthonous) picornaviruses in the lake
- Putatively allochthonous picornaviruses are negatively impacted by lower pH and higher temperature, which may indicate that they are less likely to be transmitted between infected and uninfected animals under future climate scenarios.
The study also revealed new knowledge about the taxonomic composition of bacterioplankton gene pools within lake ecosystems, which provides important background data for understanding biogeochemical cycles in the lakes. Furthermore, the study enhances our understanding of viral diversity and its interactions with climate in aquatic ecosystems, expanding the diversity of known RNA viruses, and furthering understanding of lake ecosystems generally.
Work published from this project:
Hewson I, Bistolas KSI, Button JB, Jackson EW (2018) “Occurrence and seasonal dynamics of RNA viral genotypes in three contrasting temperate lakes” PLoS One 13: e0194419

